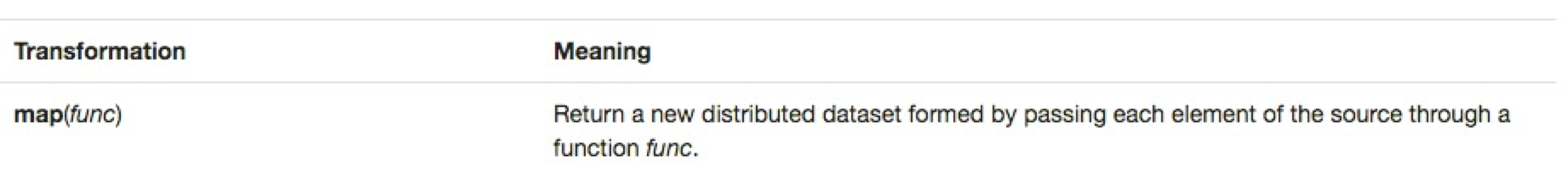
I'm writing this blog post for those learning Spark's RDD programming model and who might have heard that using the mapPartitions() transformation is usually faster than its map() brethren and wondered why. If in doubt... RTFM, which brought me to the following excerpt from http://spark.apache.org/docs/latest/programming-guide.html.
The good news is that the answer is right there. It just might not be as apparent for those from Missouri (you know... the "Show Me State") and almost all of us can benefit from a working example. If you're almost there from reading the description above then https://bzhangusc.wordpress.com/2014/06/19/optimize-map-performamce-with-mappartitions/ might just push you over the "aha" moment. It worked for me, but I wanted to build a fresh example all on my own which I'll try to document well enough for you to review and even recreate if you like.
If you made it this far then you know that Spark's Resilient Distributed Dataset (RDD) is a partitioned beast all its own somewhat similar to how HDFS breaks files up into blocks. In fact, when we load a file in Spark from HDFS by default the number of RDD partitions is the same as the number of HDFS blocks. You can set the number of partitions you want for the RDD (I'm breaking my example into three) and that itself is a topic for another, even bigger, discussion. If you are still with me then you probably know that "narrow" transformations/tasks happen independently on each of the partitions. Therefore, the well-used map() function is working in parallel on each of the RDD's partition that it is walking through.
That's good news – in fact, that's great news as these narrow tasks are key to performance! The sister mapPartitions() transformation also works independently on the partitions; so what's so special that makes it run better in most cases? Well... it comes down to the fact that map() exercises the function being utilized at a per element level while mapPartitions() exercises the function at the partition level. What does that mean in practice?
It means that if we have 100K elements in a particular RDD partition then we will fire off the function being used by the mapping transformation 100K times when we use map(). Conversely, if we use mapPartitions() then we will only call the particular function one time, but we will pass in all 100K records and get back all responses in one function call.
That means, we could get a big lift in the fact that we aren't exercising the particular function so many times, especially if the function is doing something expensive each time that it wouldn't need to do if we passed in all the elements at once.
Hmmm... if that doesn't make immediate sense, let's do some housekeeping for our example and then circle-back to this thought.
This next little bit simple loads up a timeless Dr Suess story into an RDD that then gets transformed into another RDD with a single word for each element. I'll explain more in a bit, but let's also cache this RDD into memory to aid our ultimate test later in the blog.
For my testing, I used version 2.4 of the Hortonworks Sandbox. Once you get this started up, SSH into the machine as root, but then switch to the mktg1 user who has a local linux account as well as an HDFS home directory. If you want to use a brand-new user, try my simple hadoop cluster user provisioning process (simple = w/o pam or kerberos) process. Once there, you can copy the contents of GreenEggsAndHam.txt onto your clipboard and paste it into the vi editor (just type "i" once vi starts and paste from your clipboard, then end it all by hitting ESC twice and typing ":" to get the command prompt and "wq" to write & quit the editor) and then copy that file into HDFS.
HW10653-2:~ lmartin$ ssh root@127.0.0.1 -p 2222 root@127.0.0.1's password: Last login: Thu May 19 22:04:06 2016 from 10.0.2.2 [root@sandbox ~]# su - mktg1 [mktg1@sandbox ~]$ vi GreenEggsAndHam.txt [mktg1@sandbox ~]$ hdfs dfs -put GreenEggsAndHam.txt [mktg1@sandbox ~]$ hdfs dfs -ls Found 1 items -rw-r--r-- 3 mktg1 hdfs 3478 2016-05-19 22:13 GreenEggsAndHam.txt
Then let's start-up the Spark shell. NOTE: Like in the following capture, I'm eliminating a lot of "noise" to keep us focused on the important stuff.
[mktg1@sandbox ~]$ spark-shell
Welcome to
____ __
/ __/__ ___ _____/ /__
_\ \/ _ \/ _ `/ __/ '_/
/___/ .__/\_,_/_/ /_/\_\ version 1.6.0
/_/
Using Scala version 2.10.5 (OpenJDK 64-Bit Server VM, Java 1.7.0_95)
Type in expressions to have them evaluated.
Type :help for more information.
SQL context available as sqlContext.
scala>
As described earlier, now we load up the Dr Suess story into an RDD that then gets split into another RDD of a single word per element. Let's also cache it to establish a baseline of simply reading through the RDD so that we don't introduce any additional variability in our comparison testing.
scala> val wordsRdd = sc.textFile("GreenEggsAndHam.txt", 3).flatMap(line => line.split(" "))
wordsRdd: org.apache.spark.rdd.RDD[String] = MapPartitionsRDD[2] at flatMap at <console>:27
scala> wordsRdd.persist() //mark it as cached
res0: wordsRdd.type = MapPartitionsRDD[2] at flatMap at <console>:27
scala> sc.setLogLevel("INFO") //enable timing messages
scala> wordsRdd.take(3) //trigger cache load
16/05/19 23:08:22 INFO DAGScheduler: Job 0 finished: take at <console>:30, took 0.565019 s
res2: Array[String] = Array(I, am, Daniel)
Just to verify that the cache is helping performance (was 0.565 secs above), let's hit it a few more times to see a must faster and consistent response.
scala> wordsRdd.take(3) 16/05/19 23:11:12 INFO DAGScheduler: Job 1 finished: take at <console>:30, took 0.044692 s res3: Array[String] = Array(I, am, Daniel) scala> wordsRdd.take(3) 16/05/19 23:11:14 INFO DAGScheduler: Job 2 finished: take at <console>:30, took 0.020183 s res4: Array[String] = Array(I, am, Daniel) scala> wordsRdd.take(3) 16/05/19 23:11:17 INFO DAGScheduler: Job 3 finished: take at <console>:30, took 0.033659 s res5: Array[String] = Array(I, am, Daniel)
Now that we got an order of magnitude speed improvement, and somewhat consistent response times, we are ready to stand up a test harness to prove that mapPartitions() is faster than map() when the function we are calling produces negative results when call once per record instead of once per partition. The wrapSingleWord() function is just going to add ">>" before each word in the RDD and slap on "<<" on the backside. And to really make the point, I've snuck in a bogus function that burns 10 millis of clock time for each function call which is my simulation of some arbitrary expensive operation. Just paste in the following functions separately in the shell.
def simulateExpensiveObjectCreation() {
Thread sleep 10
}
def wrapSingleWord(original: String) : String = {
simulateExpensiveObjectCreation()
val sb = new StringBuilder()
return sb.append(">>").append(original).append("<<").toString()
}
Next up, let's run our RDD through a map() transformation and get a feel of the timing it takes to exercise this model where the wrapSingleWord() function gets call for each word in the RDD.
scala> wordsRdd.map(wrapSingleWord).take(10) 16/05/20 00:18:46 INFO DAGScheduler: Job 7 finished: take at <console>:34, took 0.151721 s res9: Array[String] = Array(>>I<<, >>am<<, >>Daniel<<, >><<, >>I<<, >>am<<, >>Sam<<, >>Sam<<, >>I<<, >>am<<) scala> wordsRdd.map(wrapSingleWord).take(10) 16/05/20 00:18:52 INFO DAGScheduler: Job 8 finished: take at <console>:34, took 0.146428 s res10: Array[String] = Array(>>I<<, >>am<<, >>Daniel<<, >><<, >>I<<, >>am<<, >>Sam<<, >>Sam<<, >>I<<, >>am<<) scala> wordsRdd.map(wrapSingleWord).take(10) 16/05/20 00:18:54 INFO DAGScheduler: Job 9 finished: take at <console>:34, took 0.153416 s res11: Array[String] = Array(>>I<<, >>am<<, >>Daniel<<, >><<, >>I<<, >>am<<, >>Sam<<, >>Sam<<, >>I<<, >>am<<)
Remember, wordsRdd.map(wrapSingleWord) is shorthand for wordsRdd.map(word => wrapSingleWord(word)) since the function takes in the entire RDD element.
In this tightly constrained Sandbox cluster, we are seeing about 0.15 secs to execute this completely. Now, to get mapPartitions() to work we need another function that does the same thing, but it has to allow the whole collection of elements to be passed into the function and it needs to return a whole collection on the way back. The following is just a jazzed up version of the earlier function.
import java.util._
import scala.collection.JavaConversions._
def wrapMultiWords(words: Iterator[String]) : Iterator[String] = {
simulateExpensiveObjectCreation()
val sb = new StringBuilder()
val wList = new ArrayList[String]()
while( words.hasNext ) {
sb.setLength(0)
wList.add( sb.append(">>").append(words.next()).append("<<").toString() )
}
return wList.iterator()
}
After you get that added into the shell we can exercise it to see how it compares.
scala> wordsRdd.mapPartitions(word => wrapMultiWords(word)).take(10) 16/05/20 00:25:51 INFO DAGScheduler: Job 10 finished: take at <console>:40, took 0.069876 s res12: Array[String] = Array(>>I<<, >>am<<, >>Daniel<<, >><<, >>I<<, >>am<<, >>Sam<<, >>Sam<<, >>I<<, >>am<<) scala> wordsRdd.mapPartitions(word => wrapMultiWords(word)).take(10) 16/05/20 00:26:04 INFO DAGScheduler: Job 11 finished: take at <console>:40, took 0.050593 s res13: Array[String] = Array(>>I<<, >>am<<, >>Daniel<<, >><<, >>I<<, >>am<<, >>Sam<<, >>Sam<<, >>I<<, >>am<<) scala> wordsRdd.mapPartitions(word => wrapMultiWords(word)).take(10) 16/05/20 00:26:07 INFO DAGScheduler: Job 12 finished: take at <console>:40, took 0.055201 s res14: Array[String] = Array(>>I<<, >>am<<, >>Daniel<<, >><<, >>I<<, >>am<<, >>Sam<<, >>Sam<<, >>I<<, >>am<<)
As you can see, the new RDD has the same data, but the performance was dramatically better finishing in about a third of the time in this testing scenario. Obviously, you'll need to do some testing with your own data and the functions that are being used in the mapping transformations, but if you do have any measurable difference in calling the related function, it will surely surface when you have to call it over and over again. While Spark is not necessarily Hadoop, the need to POC your particular problem and validate your hypothesis is just as important in this space. With that, you will likely see improvements by moving from map() to mapPartitions() when your related function can process all the elements at once.